Jen Cook-Chrysos/Whitehead Institute
Blending machine learning and biology to predict cell fates and other changes
Imagine a ball thrown in the air: it curves up, then down, tracing an arc to a point on the ground some distance away. The path of the ball can be described with a simple mathematical equation, and if you know the equation, you can figure out where the ball is going to land. Biological systems tend to be harder to forecast, but Whitehead Institute Member Jonathan Weissman, postdoc in his lab Xiaojie Qiu, and collaborators at the University of Pittsburgh School of Medicine are working on making the path taken by cells as predictable as the arc of a ball. Rather than looking at how cells move through space, they are considering how cells change with time.
Weissman, Qiu, and collaborators Jianhua Xing, professor of computational and systems biology at the University of Pittsburgh School of Medicine, and Xing lab graduate student Yan Zhang have built a machine learning framework that can define the mathematical equations describing a cell’s trajectory from one state to another, such as its development from a stem cell into one of several different types of mature cell. The framework, called dynamo, can also be used to figure out the underlying mechanisms—the specific cocktail of gene activity—driving changes in the cell. Researchers could potentially use these insights to manipulate cells into taking one path instead of another, a common goal in biomedical research and regenerative medicine.
The researchers describe dynamo in a paper published in the journal Cell on February 1. They explain the framework’s many analytical capabilities and use it to help understand mechanisms of human blood cell production, such as why one type of blood cell forms first (appears more rapidly than others).
“Our goal is to move towards a more quantitative version of single cell biology,” Qiu says. “We want to be able to map how a cell changes in relation to the interplay of regulatory genes as accurately as an astronomer can chart a planet’s movement in relation to gravity, and then we want to understand and be able to control those changes.”
How to map a cell’s future journey
Dynamo uses data from many individual cells to come up with its equations. The main information that it requires is how the expression of different genes in a cell changes from moment to moment. The researchers estimate this by looking at changes in the amount of RNA over time, because RNA is a measurable product of gene expression. In the same way that knowing the starting position and velocity of a ball is necessary to understand the arc it will follow, researchers use the starting levels of RNAs and how those RNA levels are changing to predict the path of the cell. However, calculating changes in the amount of RNA from single cell sequencing data is challenging, because sequencing only measures RNA once. Researchers must then use clues like RNA-being-made at the time of sequencing and equations for RNA turnover to estimate how RNA levels were changing. Qiu and colleagues had to improve on previous methods in several ways in order to get clean enough measurements for dynamo to work. In particular, they used a recently developed experimental method that tags new RNA to distinguish it from old RNA, and combined this with sophisticated mathematical modeling, to overcome limitations of older estimation approaches.
The researchers’ next challenge was to move from observing cells at discrete points in time to a continuous picture of how cells change. The difference is like switching from a map showing only landmarks to a map that shows the uninterrupted landscape, making it possible to trace the paths between landmarks. Led by Qiu and Zhang, the group used machine learning to reveal continuous functions that define these spaces.
“There have been tremendous advances in methods for broadly profiling transcriptomes and other ‘omic’ information with single-cell resolution. The analytical tools for exploring these data, however, to date have been descriptive instead of predictive. With a continuous function, you can start to do things that weren’t possible with just accurately sampled cells at different states. For example, you can ask: if I changed one transcription factor, how is it going to change the expression of the other genes?” says Weissman, who is also a professor of biology at the Massachusetts Institute of Technology (MIT), a member of the Koch Institute for Integrative Biology Research at MIT, and an investigator of the Howard Hughes Medical Institute.
Dynamo can visualize these functions by turning them into math-based maps. The terrain of each map is determined by factors like the relative expression of key genes. A cell’s starting place on the map is determined by its current gene expression dynamics. Once you know where the cell starts, you can trace the path from that spot to find out where the cell will end up.
Xiaojie Qiu/Whitehead Institute
The researchers confirmed dynamo’s cell fate predictions by testing it against cloned cells–cells that share the same genetics and ancestry. One of two nearly-identical clones would be sequenced while the other clone went on to differentiate. Dynamo’s predictions for what would have happened to each sequenced cell matched what happened to its clone.
Moving from math to biological insight and non-trivial predictions
With a continuous function for a cell’s path over time determined, dynamo can then gain insights into the underlying biological mechanisms. Calculating derivatives of the function provides a wealth of information, for example by allowing researchers to determine the functional relationships between genes—whether and how they regulate each other. Calculating acceleration can show that a gene’s expression is growing or shrinking quickly even when its current level is low, and can be used to reveal which genes play key roles in determining a cell’s fate very early in the cell’s trajectory. The researchers tested their tools on blood cells, which have a large and branching differentiation tree. Together with blood cell expert Vijay Sankaran of Boston Children’s Hospital, the Dana-Farber Cancer Institute, Harvard Medical School, and Broad Institute of MIT and Harvard, and Eric Lander of Broad Institute, they found that dynamo accurately mapped blood cell differentiation and confirmed a recent finding that one type of blood cell, megakaryocytes, forms earlier than others. Dynamo also discovered the mechanism behind this early differentiation: the gene that drives megakaryocyte differentiation, FLI1, can self-activate, and because of this is present at relatively high levels early on in progenitor cells. This predisposes the progenitors to differentiate into megakaryocytes first.
The researchers hope that dynamo could not only help them understand how cells transition from one state to another, but also guide researchers in controlling this. To this end, dynamo includes tools to simulate how cells will change based on different manipulations, and a method to find the most efficient path from one cell state to another. These tools provide a powerful framework for researchers to predict how to optimally reprogram any cell type to another, a fundamental challenge in stem cell biology and regenerative medicine, as well as to generate hypotheses of how other genetic changes will alter cells’ fate. There are a variety of possible applications.
“If we devise a set of equations that can describe how genes within a cell regulate each other, we can computationally describe how to transform terminally differentiated cells into stem cells, or predict how a cancer cell may respond to various combinations of drugs that would be impractical to test experimentally,” Xing says.
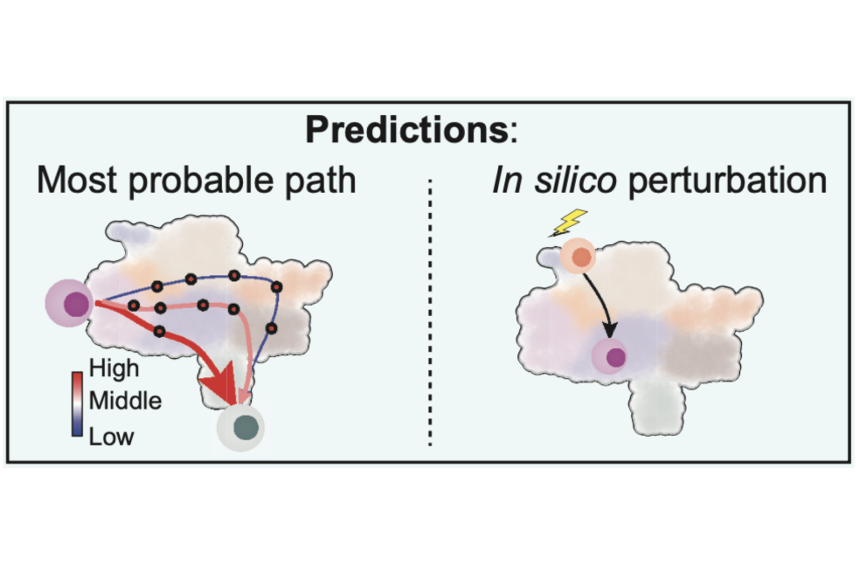
Dynamo’s computational modeling can be used to predict the most likely path that a cell will follow when reprogramming one cell type to another, as well as the path that a cell will take after specific genetic manipulations.
Xiaojie Qiu/Whitehead Institute
Dynamo moves beyond merely descriptive and statistical analyses of single cell sequencing data to derive a predictive theory of cell fate transitions. The dynamo toolset can provide deep insights into how cells change over time, hopefully making cells’ trajectories as predictable for researchers as the arc of a ball, and therefore also as easy to change as switching up a pitch.
Xiaojie Qiu, Yan Zhang, Jorge D. Martin-Rufino, Chen Weng, Shayan Hosseinzadeh, Dian Yang1,2, Angela N. Pogson, Marco Y. Hein, Kyung Hoi (Joseph) Min, Li Wang, Emanuelle I. Grody, Matthew J. Shurtleff, Ruoshi Yuan, Song Xu, Yian Ma, Joseph M. Replogle, Eric S. Lander, Spyros Darmanis, Ivet Bahar, Vijay G. Sankaran, Jianhua Xing, Jonathan S Weissman. “Mapping Transcriptomic Vector Fields of Single Cells.” Cell, Feb 1. DOI: https://doi.org/10.1016/j.cell.2021.12.045
Contact
Communications and Public Affairs
Phone: 617-452-4630
Email: newsroom@wi.mit.edu