
A "model" parasite
Whitehead Institute researchers unravel the unique biology of apicomplexans — the parasites responsible for malaria, toxoplasmosis, and other diseases impacting global health.
Apicomplexa: A brood of parasites
Malaria, cryptosporidiosis, and toxoplasmosis affect millions of people each year, killing an estimated 600,000 annually, mostly children under five in developing countries. Billions of dollars are spent each year to control and eliminate these diseases, according to the World Health Organization (WHO). Each of these diseases is caused by a different apicomplexan, a group of parasites that infect almost all animal species.
Toxoplasma gondii (T. gondii), which causes the disease toxoplasmosis, has a unique physiology that has allowed it to parasitize its hosts, yet it retains many features in common with other apicomplexans. Using T. gondii as a “model parasite”, Whitehead Member Sebastian Lourido is deciphering apicomplexans’ unique biology and uncovering aspects that could be harnessed to disrupt the parasites’ ability to proliferate and infect their hosts.
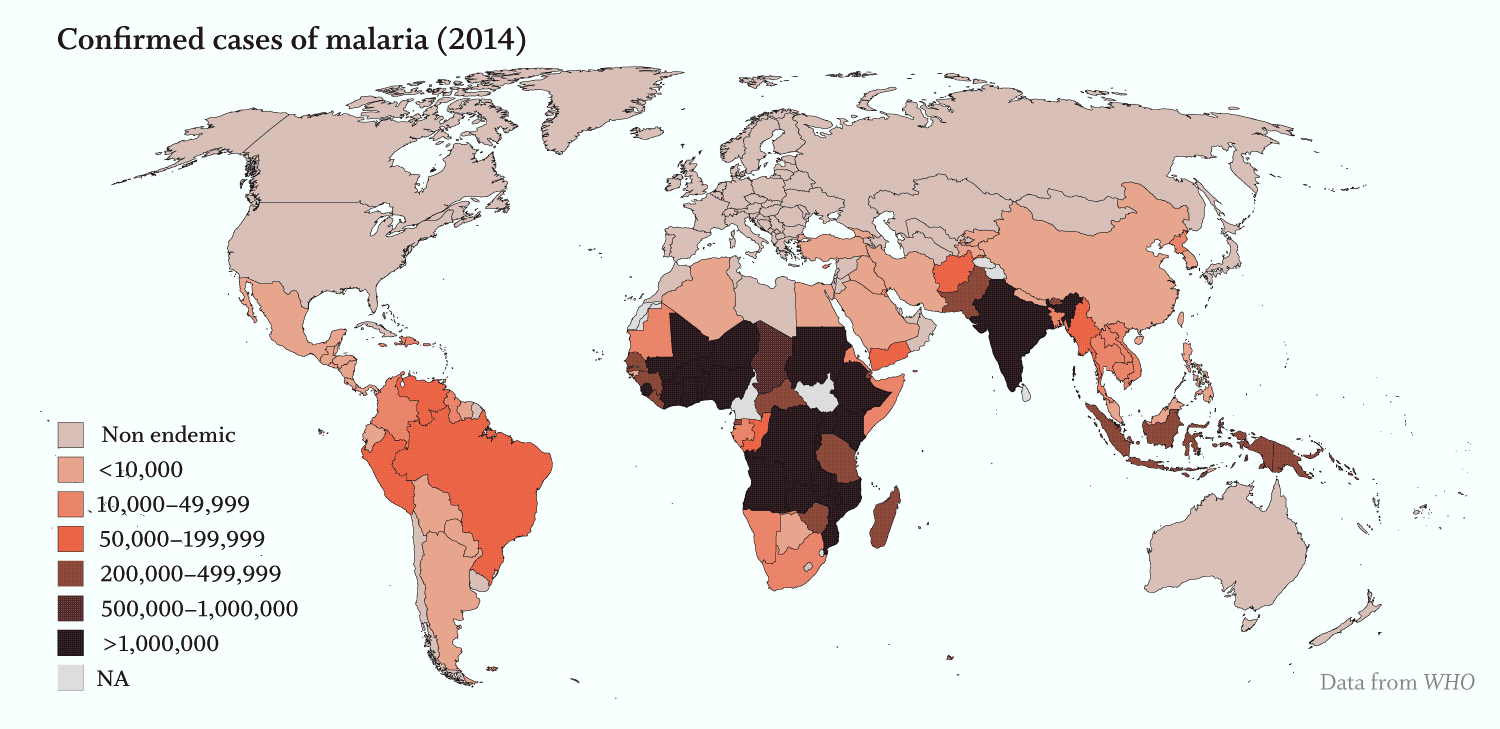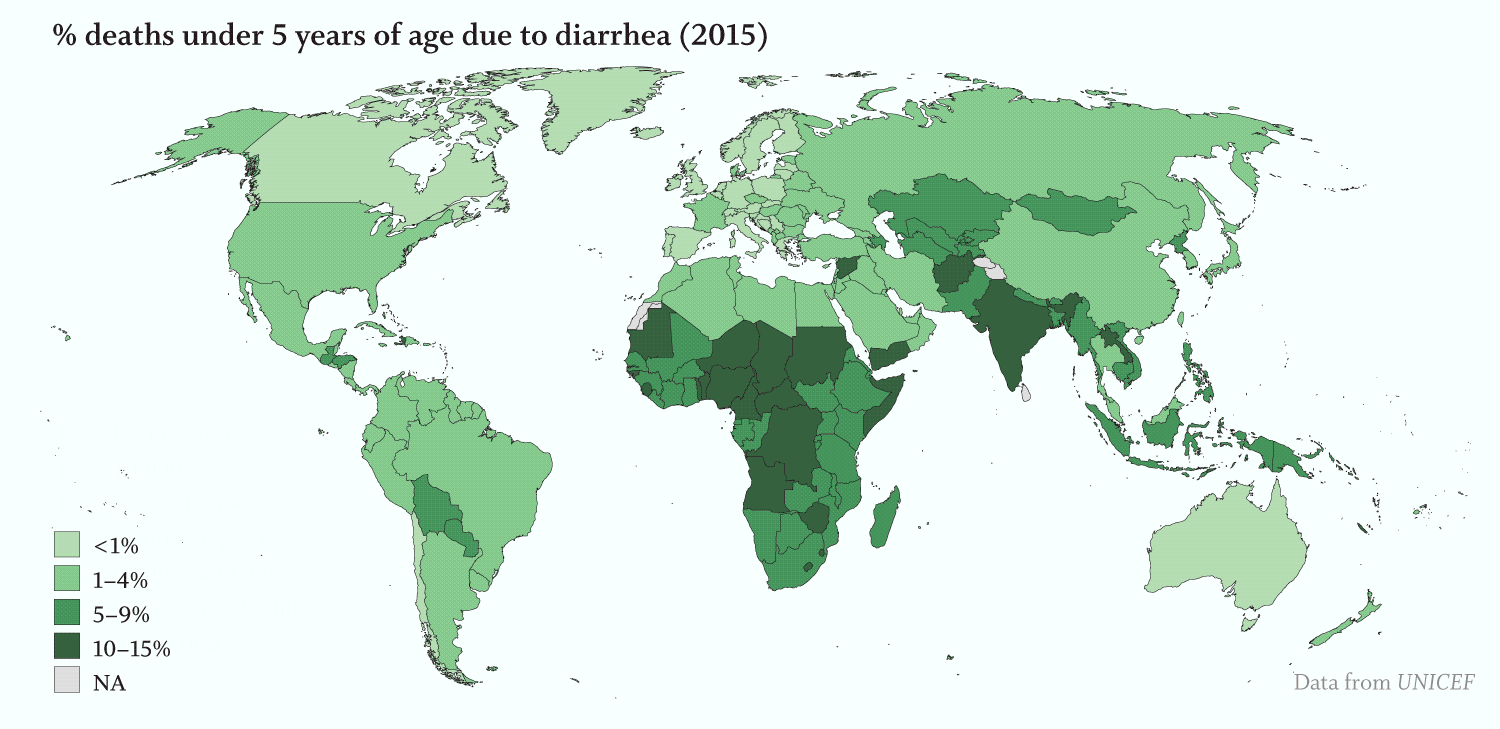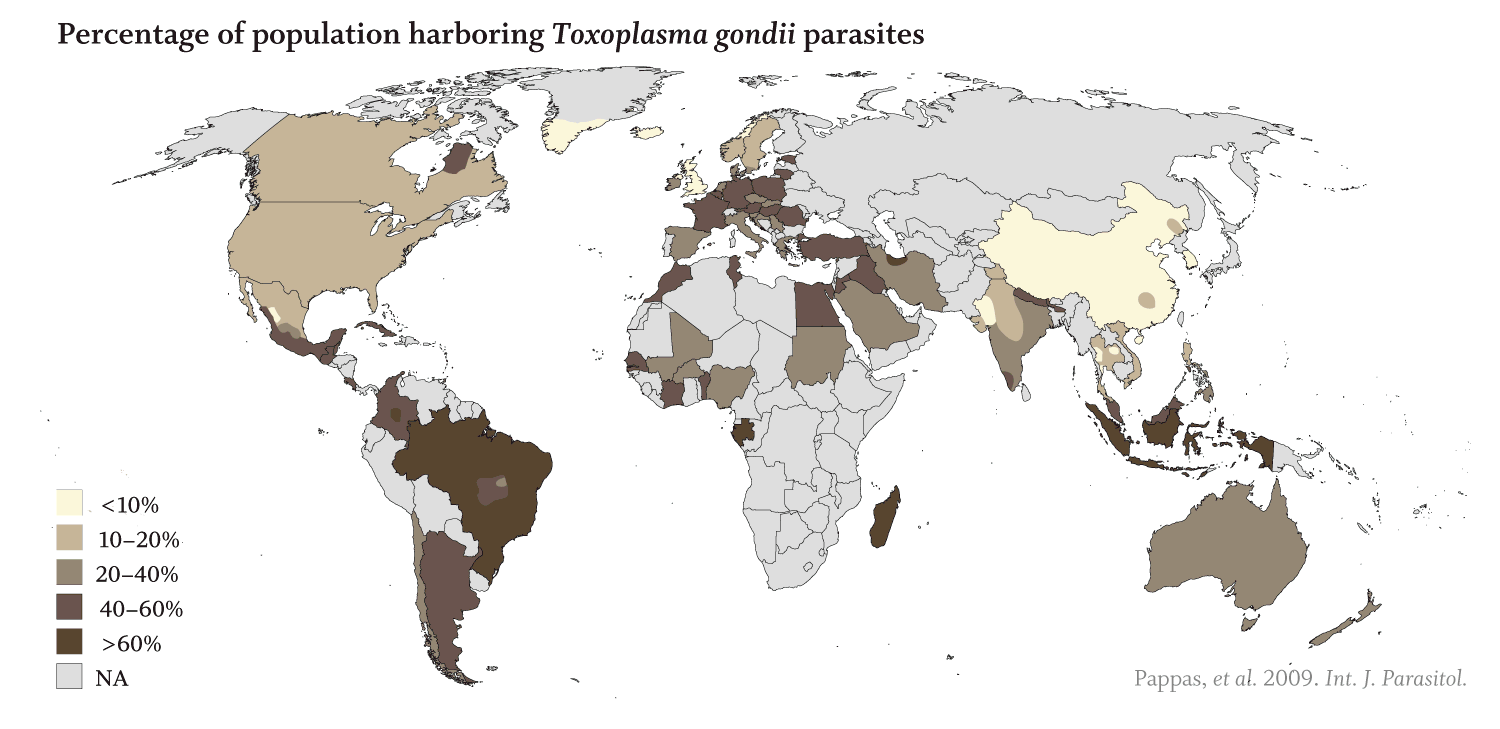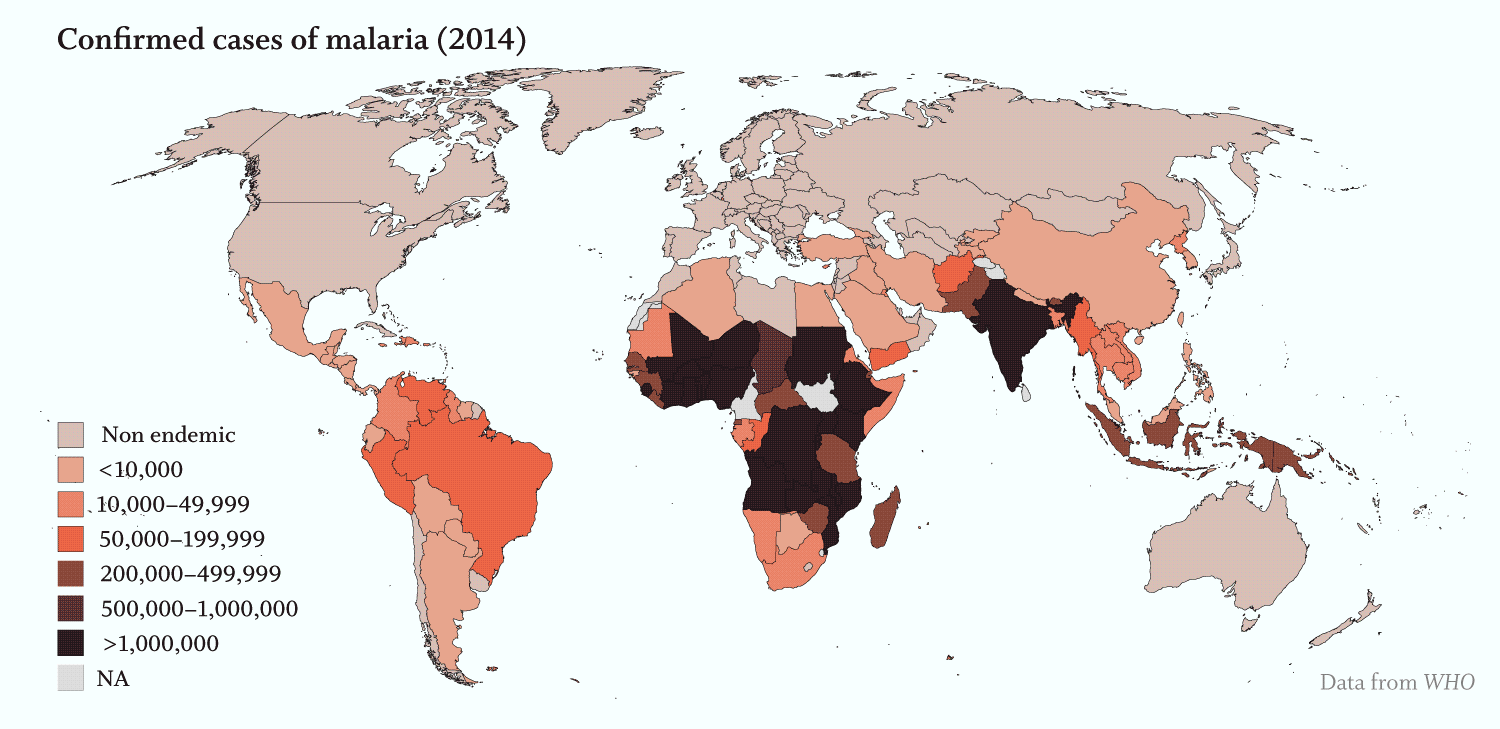
Apicomplexans’ toll on humans is staggering:
· Malaria, caused by several Plasmodium species of Apicomplexa, was responsible for over 200 million infections and more than 400,000 deaths, primarily in young children, in 2017 (WHO).
· Severe diarrhea kills an estimated 525,000 children under five each year (WHO). Over 200,000 of those deaths can be attributed to cryptosporidiosis, which is caused by the species of the apicomplexan Cryptosporidium (Sow et al., 2016, PLoS Negl Trop Dis.).
· 25% of the global population is infected with T. gondii with rates reaching over 60% in some areas (Pappas et al. 2009, Int. J. Parasitol.). Toxoplasmosis can cause an array of serious neurological disorders in those with weakened immune systems and can be lethal or lead to birth defects in a developing fetus. In an estimated 2% of infected individuals, toxoplasmosis causes retinal lesions (Holland, 2003, Am J Ophthalmol.).
A parasitic relationship, separated by a billion years of evolution

The diagram above depicts the evolutionary relationships between organisms — species separated by many branches are more distantly related than those divided by fewer branches. Apicomplexans and humans are separated by multiple branches and more than a billion years of evolution. In fact, apicomplexans are actually more closely related to plants than animals, having evolved from a non-parasitic ancestor about 700 to 900 million years ago that, like green plants, used photosynthesis to generate energy from sunlight. So far, scientists have studied only about half of the known apicomplexan genes, leaving the rest of their 8,000 predicted protein-coding genes uncharacterized.
Although key genes important for fundamental processes have remained fairly stable over the billion years since apicomplexans and their hosts diverged, other parts of the apicomplexan genomes evolved as they adapted to a parasitic lifestyle. The genes that emerged as unique to apicomplexans, such as those encoding factors involved in entering or exiting host cells, potentially represent therapeutic targets because curtailing their expression could hamstring — or even eliminate — the parasites without harming the host.
Analyzing a unique biology
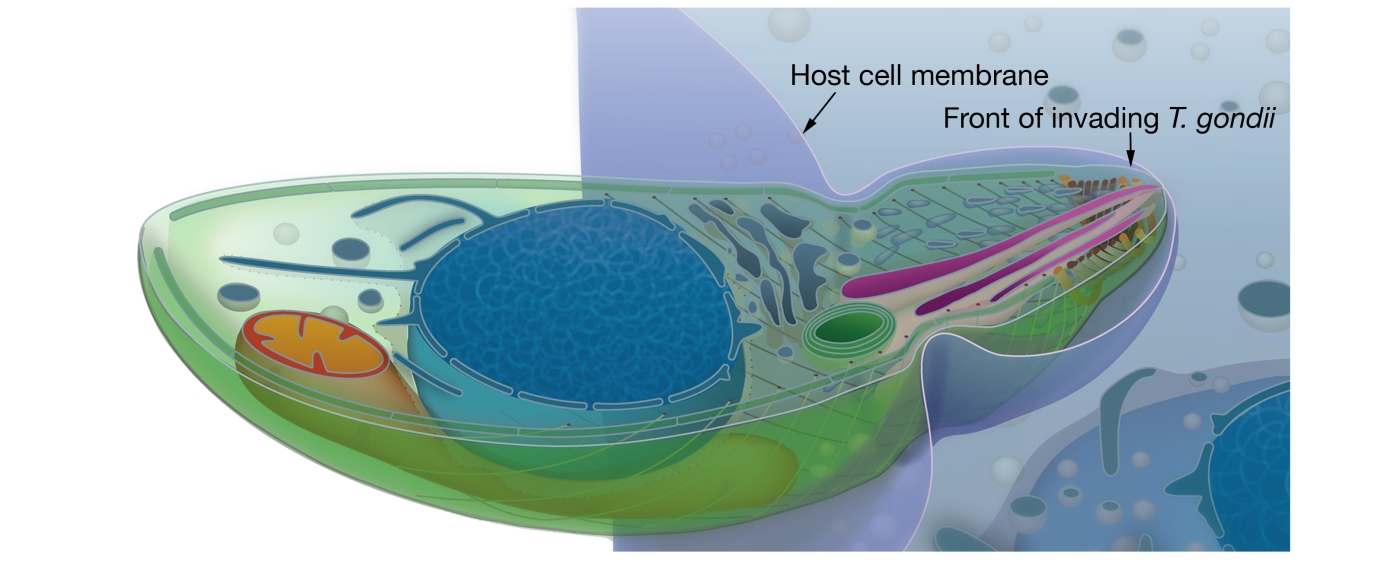
Understanding apicomplexans and their distinct biology has been challenging at least in part because the tools — genomic analysis, genetic engineering, and culture systems — that scientists use to study and understand more traditional model organisms in the lab, such as mice, are difficult to apply in apicomplexans. Moreover, apicomplexans may spend different stages of their lives in different hosts, so studying a parasite’s complete life cycle may require studying and culturing multiple organisms or their tissues. For example, Plasmodium falciparum, which causes malaria, spends part of its life cycle in mosquitoes and another in humans.

Whitehead Institute Member Sebastian Lourido (above) and his lab study the parasite T. gondii, which causes toxoplasmosis.
Gretchen Ertl/Whitehead Institute
Unlike the Plasmodium parasites that cause malaria, T. gondii is relatively easy to culture in the lab. Lourido, who is also an assistant professor of biology at Massachusetts Institute of Technology (MIT), and his lab are using this organism to unravel many elemental questions about apicomplexans: How do they infect their host cells? What do they require to reproduce? How do they break out of their host cell to infect more cells?
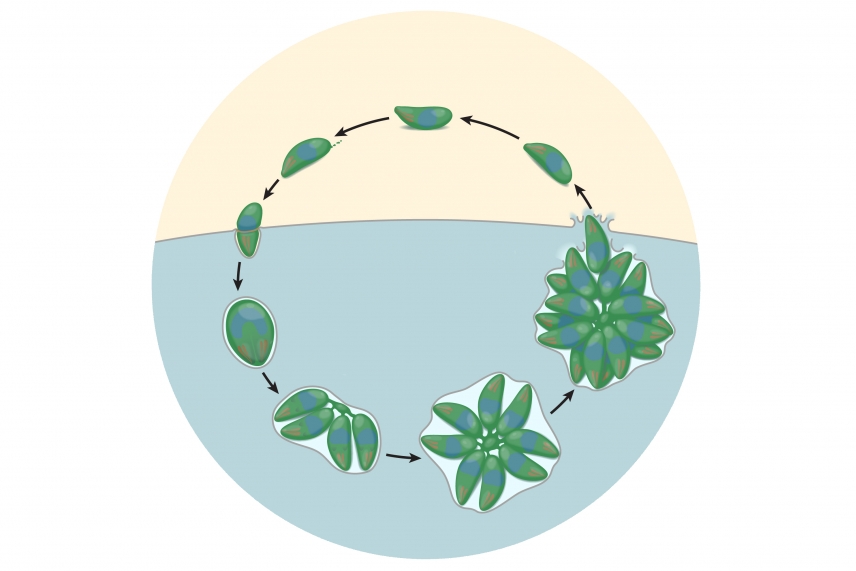
During its lifecycle in human host cells, T. gondii invades a host (blue), reproduces and creates characteristic rosette clusters of parasites, then breaks out of the cell before invading another host cell.
Adapted CRISPR/Cas9 gene editing system reveals first genome-wide glimpse of apicomplexan genomic profile

T. gondii parasites (magenta) surrounded by a web of their human cell’s cytoskeleton (white).
Courtesy of Clare Harding/Whitehead Institute
Researchers in Lourido’s lab are working to decipher the 50% of the T. gondii’s genome that remains to be characterized. To do so, they adapted the CRISPR/Cas9 gene editing system to work in T. gondii. Using CRISPR/Cas9, researchers can cut T. gondii’s DNA at specific sites to disable particular genes. With this approach, they were able to efficiently conduct genome-wide screens to identify genes that are functionally important to the parasite.
For this screen the Lourido lab used their adapted CRISPR/Cas9 gene editing system to remove the function — one at a time — of each of T. gondii’s ~8,000 protein-coding genes. The resulting altered parasites were then cultured with human host cells. After a period of time, the scientists tallied the number of parasites present with each modification to assess how disabling a particular gene’s function affects the parasites’ reproduction and survival. Altered parasites that successfully proliferated despite missing a gene’s function were deemed to have alterations in a gene that is dispensable, whereas modified parasites that did not thrive were deemed to have alterations in genes that are important for fitness.
Screen identifies apixomplexan-specific proteins
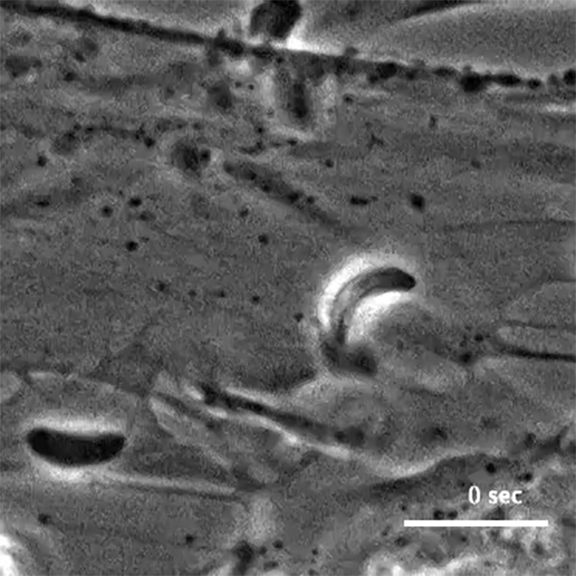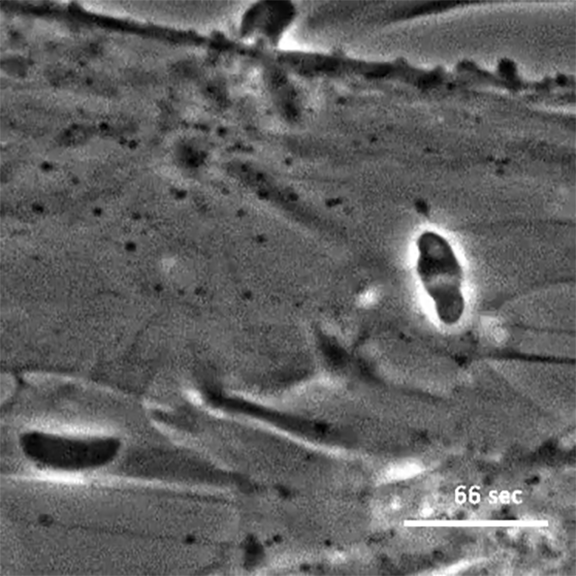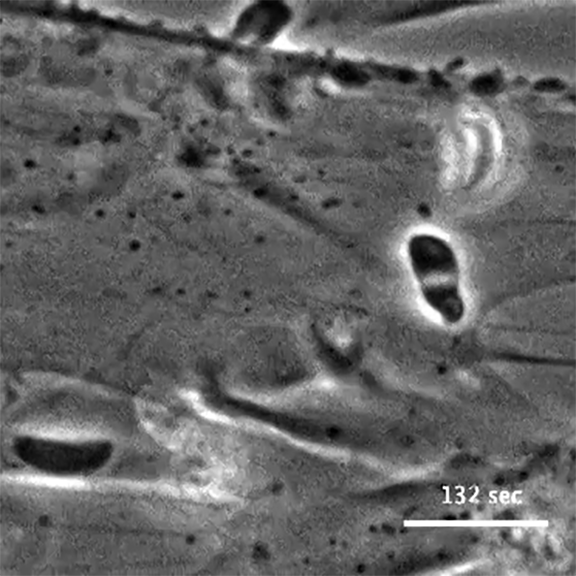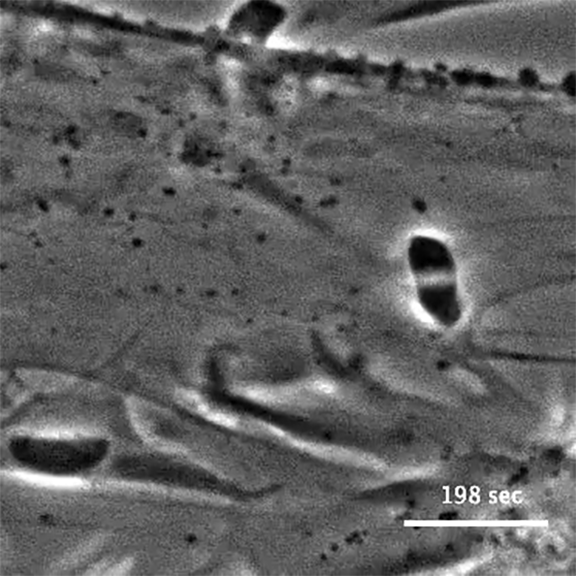
The initial screen of the T. gondii genome, led by Lourido lab research assistant Saima Sidik and postdoctoral researcher Diego Huet, identified a number of genes that encode indispensable conserved apicomplexan proteins, called ICAPs for short (Sidik et al. 2016, Cell). One ICAP identified by Sidik and Huet is an invasion factor called the claudin-like apicomplexan microneme protein (CLAMP).

In the same Cell paper, researchers described how they determined that CLAMP is critical to the initiation of host cell invasion by T. gondii. Working with Jacquin Niles and members of his lab in the MIT Department of Biological Engineering, the Lourido team also showed that CLAMP is required for the parasites that cause malaria to survive when grown in red blood cells.
Tapping the energy source

Whitehead Institute Member Sebastian Lourido (right) with postdoctoral researcher Diego Huet (left) and research associate Saima Sidik (center) from his lab.
Clare Harding/Whitehead Institute
In a recent paper published in the journal eLife, Lourido and first author Huet, identified a protein in apicomplexans that is crucial for creating adenosine triphosphate (ATP), the universal energy storage unit of cells. ATP is essential for cells’ survival and without it, cellular processes would stall. Most organisms have an enzyme, called ATP synthase, that creates ATP by converting the energy of a proton gradient across a membrane into mechanical energy. As the protons move through the ATP synthase, it spins like a turbine. This movement powers the formation of ATP. For the enzyme to work properly, a portion of the ATP synthase acts as a scaffold, or stator, by counteracting the rotation of the turbine-like part of the enzyme.
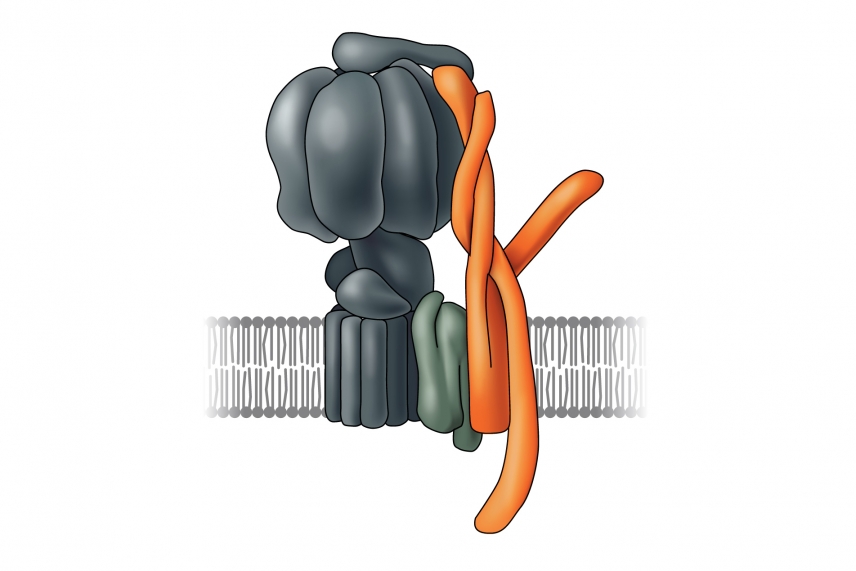
In T. gondii’s ATP synthase, the stator, which was identified by Lourido and Huet, is shown in orange.
Although most components of the ATP synthase are conserved between apicomplexans and humans, scientists had been unable to pinpoint the gene encoding the essential stator portion – no DNA sequence in the apicomplexan genome resembles the sequences of known stator genes. Using a genomic approach, Huet and Lourido analyzed the predicted function and structure of the ICAPs present in the mitochondrion, the ATP synthase’s home in all organisms. To their surprise, the predicted shape of one of the ICAPs resembles a stator subunit found in yeast and mammals.
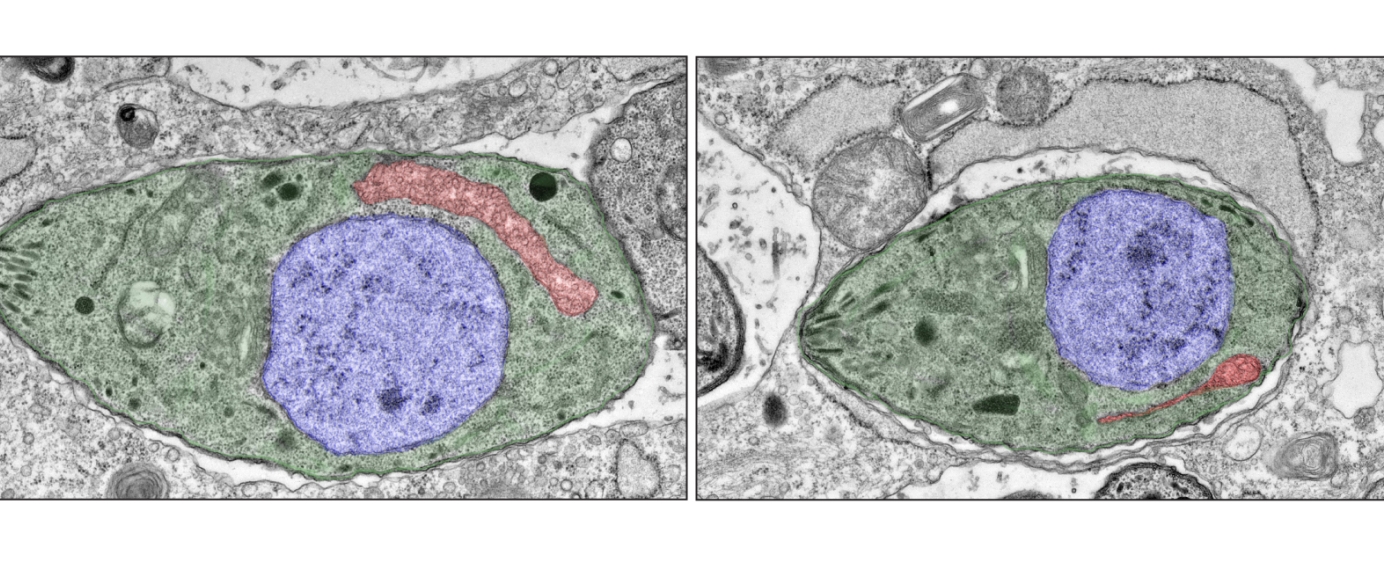
Electron microscopy images of one individual wild-type T. gondii parasite (green, left) and one individual T. gondii parasite with an incomplete ATP synthase (green, right) showing that the mitochondrion (red) is severely reduced when the assembly of the ATP synthase is impaired. For clarity, the nucleus is highlighted in blue.
Diego Huet/Whitehead Institute
When Huet and Lourido mutated or removed the stator subunit, the parasite’s ATP synthase failed to function properly, damaging the structure and performance of its mitochondria and halting the the parasite’s growth.
The beginning of a parasitic relationship

T. gondii parasites (yellow) surrounded by a network of human host cell’s cytoskeleton (red) and DNA (blue).
Courtesy of Clare Harding/Whitehead Institute
For Lourido and his lab, T. gondii’s unique stator protein is just one example of how these extraordinary apicomplexan organisms have evolved and adapted. By tailoring current tools and inventing new ones, Lourido’s investigations into T. gondii’s biology have the potential to reveal important insights into this family of parasites that impacts millions of people each year.
Credits & References
Credits:
Written and produced by Nicole Giese Rura and Whitehead Institute
Illustrations and animations by Andrew Tubelli
Cover image courtesy of Clare Harding/Whitehead Institute
Special thanks to Sebastian Lourido and his lab, especially Clare Harding, Diego Huet, and Saima Sidik
References:
WHO: Global Health Observatory (GHO data) for number of malaria cases
UNICEF: Diarrhoea as a cause of death in children under 5
Holland GN. (2003) Ocular toxoplasmosis: a global reassessment. Part I: epidemiology and course of disease. Am J Ophthalmol. Dec;136(6): 973-988. https://doi.org/10.1016/j.ajo.2003.09.040
Huet D, Rajendran E, van Dooren GG, Lourido S. (2018) Identification of Cryptic Subunits from an Apicomplexan ATP Synthase. eLife. 2018;7:e38097. https://doi:10.7554/eLife.38097
Pappas G, Roussos N, Falagas ME. (2009) Toxoplasmosis Snapshots: Global Status of Toxoplasma gondii Seroprevalence and Implications for Pregnancy and Congenital Toxoplasmosis. Int J Parasitol. Oct;39(12):1385-94. https://doi: 10.1016/j.ijpara.2009.04.003
Sidik SM, Huet D, Ganesan SM, Huynh MH, Wang T, Nasamu AS, Thiru P, Saeij JPJ, Carruthers VB, Niles JC, Lourido S. (2016) A Genome-wide CRISPR Screen in Toxoplasma Identifies Essential Apicomplexan Genes. Cell. Sep 8;166(6):1423-1435.e12. https://doi:10.1016/j.cell.2016.08.019
Sow SO, Muhsen K, Nasrin D, Blackwelder WC, Wu Y, Farag TH, et al. (2016) The Burden of Cryptosporidium Diarrheal Disease among Children < 24 Months of Age in Moderate/High Mortality Regions of Sub-Saharan Africa and South Asia, Utilizing Data from the Global Enteric Multicenter Study (GEMS). PLoS Negl Trop Dis. 10(5): e0004729. https://doi.org/10.1371/journal.pntd.0004729
Yahata K, Treeck M, Culleton R, Gilberger T-W, Kaneko O (2012) Time-Lapse Imaging of Red Blood Cell Invasion by the Rodent Malaria Parasite Plasmodium yoelii. PLoS ONE. 7(12): e50780. https://doi.org/10.1371/journal.pone.0050780
Contact
Communications and Public Affairs
Phone: 617-452-4630
Email: newsroom@wi.mit.edu
