Scientists identify gene required for differentiation of breast stem cells

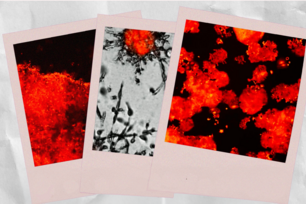
The gene RUNX1 is required for human breast stem cells to differentiate. In the slide on the left, RUNX1 is inhibited in undifferentiated primary human mammary stem cells. When RUNX1 is turned on, the stem cells differentiate into basal (red) and luminal (green) cells after 96 hours.
Courtesy of PLOS Computational Biology
CAMBRIDGE, Mass. – A novel analytical method has enabled Whitehead Institute scientists to identify a regulatory gene required for breast stem cells to differentiate. Inhibition of this gene, known as RUNX1, prevents cells from exiting the stem cell state.
“We’re very excited about this finding, as it suggests a new solution to the problem of expanding stem cells in culture,” says Whitehead Member Piyush Gupta. “Rather than trying to recreate stem cell niches in the lab, which is extremely challenging, one could simply trap stem cells in a bipotent state by inhibiting genes required for them to differentiate, such as RUNX1. This alternative approach opens up a world of possibilities for regenerative medicine.”
Until now, the search for genes controlling stem cell self-renewal and differentiation has relied on markers that label specific cell states. When available, such markers have allowed researchers to track the stem cells and observe their behavior. The problem, however, is that very few markers are available to distinguish stem cells from their slightly more differentiated counterparts, known as progenitors.
The Gupta lab devised a method called perturbation-expression analysis of cell states, or PEACS, that identifies genes regulating cell state by tracking the changes in a mixture of stem cells, early progenitors, and more differentiated cell types as they respond to factors that affect the cells’ stemness. PEACS is described in the current issue of PLOS Computational Biology.
To test PEACS’ capabilities, Ethan Sokol, a graduate student in Gupta’s lab, analyzed a line of human breast stem cells. Breast tissue is composed of several types of tissue, such as lobules that produce milk and the ducts that transport it. During development, these tissues are formed by mammary stem cells.
PEACS identified three genes in the cells that cause significant changes in the ratio of stem cells to differentiated cells. Two of the factors have established roles in mammary stem cell regulation and differentiation. The third, RUNX1, was not a known regulator of mammary stem cells but has been shown to be deregulated or mutated in some leukemias and breast cancers. Whitehead Member Peter Reddien also recently determined that a RUNX1 homolog active in flatworm stem cells is vital to regeneration in response to wounding.
To observe RUNX1’s role in breast stem cells, Sokol inhibited the gene, trapping the cells in a state of stasis and causing them to form balls of undifferentiated stem cells. When RUNX1 is expressed again, the spheres of cells sprout ducts and lobules comprising differentiated cells.
Because RUNX1 plays an important role in mammary stem cells and in breast cancer, Gupta would like to know more about the genes it regulates and the upstream factors that control its activation.
As for PEACS, Sokol wants to see if it can be used to help find drugs that control the cell state of stem cells.
“We think it could be really interesting to look for chemical inhibitors or activators of self-renewal in different stem cell populations,” says Sokol. “Using our algorithm, you could screen a chemical library and find which ones affect the cell state, either differentiation or self-renewal. Those could be interesting candidates for either regenerative medicine or maintaining a population of cells in a pure state for regenerative medicine.”
The PEACS code is available at http://guptalab.wi.mit.edu.
* * *
Piyush Gupta’s primary affiliation is with Whitehead Institute for Biomedical Research, where his laboratory is located and all his research is conducted. He is also an assistant professor of biology at Massachusetts Institute of Technology.
* * *
Citation
Sokol, E. S., Sanduja, S., Jin, D. X., Miller, D. H., Mathis, R. A., & Gupta, P. B. (2015). Perturbation-expression analysis identifies RUNX1 as a regulator of human mammary stem cell differentiation. PLoS Comput Biol, 11(4), e1004161.
Topics
Contact
Communications and Public Affairs
Phone: 617-452-4630
Email: newsroom@wi.mit.edu