Genetic master controls expose cancers’ Achilles’ heel
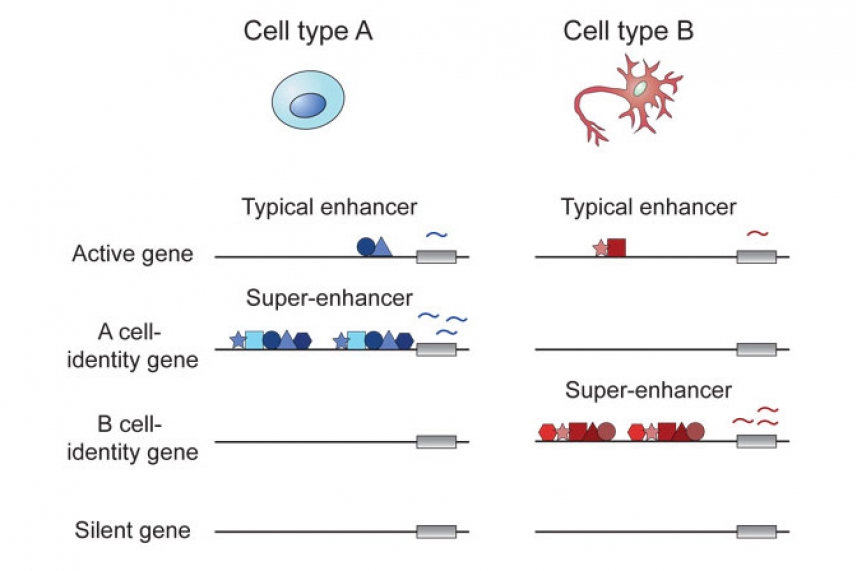
Diagram comparing enhancers and super-enhancers of two cell types
Courtesy of Cell Press
CAMBRIDGE, Mass.– In a surprising finding that helps explain fundamental behaviors of normal and diseased cells, Whitehead Institute scientists have discovered a set of powerful gene regulators dubbed “super-enhancers” that control cell state and identity. Healthy cells employ these super-enhancers to control genes responsible for cellular functions and developmental transitions—such as that from embryonic stem cell to nerve cell—but cancer cells are able to assemble their own insidious super-enhancers to overproduce harmful oncogenes that lead to aggressive tumors.
“We have been marveling at the complexity of cellular control, with millions of enhancers controlling tens of thousands of genes in the vast array of cells that comprise human beings,” says Whitehead Member Richard Young. “So it was a surprise to find that only a few hundred super-enhancers control most key genes that give each cell its special properties and functions, and furthermore, that these special controls are hijacked in cancer and other diseases.”
The findings are described in dual papers from Young and collaborators at Dana-Farber Cancer Institute published together in the April 11 edition of the journal Cell.
In the first work, the Young lab establishes a model of gene regulation in normal cells that appears to be dramatically less complex and more solvable than previously thought. To date, a vast body of research—including that of the recently described ENCODE (Encyclopedia of DNA Elements) project—has identified more than one million enhancers or “switches” that control gene expression in mammalian cells. Deciphering the precise function and target gene for each of these switches will be a daunting task, but Young and colleagues have found something of a shortcut to solving the core gene control circuitry. They show that only a few hundred special switches—that is, super-enhancers—control the key genes that actually make each cell different.
“What is fantastic about this concept is its simplicity,” notes Denes Hnisz, a Young lab postdoctoral scientist and a co-author of the first Cell paper. “We found that genes that are especially important for each cell are regulated by these specialized enhancers. But we also discovered that the super-enhancers are especially quick to change during development, and thus loss of old super-enhancers and establishment of new ones drives cell identity changes during development.”
Young says such changes in cell identity probably begin and end with the super-enhancers, which, though powerful, are also exquisitely sensitive to alterations in their environment. In fact, as differentiation begins, active super-enhancers are decommissioned, leading to changes in gene expression programs that fall under the control of newly established super-enhancers. It’s a process that adds remarkable insight to our understanding of how a fertilized egg eventually gives rise to the more than one trillion cells of the human body.
“The discovery of super-enhancers promises to help us solve the regulatory circuitry of all human cells,” Young says. That includes cancer cells.
While mapping the locations of super-enhancers along the genome of multiple myeloma (MM) cells, which are especially aggressive blood cancer cells, Young lab scientists found them in areas associated with known cancer-causing genes, including the notorious MYC oncogene. It turns out these MM cells were forming their own super-enhancers to drive dangerous overexpression of their oncogenes. Moreover, this phenomenon was not limited to MM cells, as the researchers identified super-enhancers at key tumor genes in small-cell lung cancer and the brain cancer glioblastoma multiforme.
Having observed how sensitive super-enhancers are to disruptions in their surroundings, Young and colleagues hypothesized that this sensitivity might represent a vulnerability in cancer cells whose oncogenes were in overdrive. Young collaborated with James Bradner, whose lab at Dana-Farber Cancer Institute developed an experimental drug, JQ1, which had been shown to selectively inhibit MYC expression in MM cells. Exactly how JQ1 was suppressing MYC had been unclear, until Young and Bradner discovered that its inhibition of a key enhancer component was especially profound at the MYC super-enhancer.
“It’s difficult not to be excited about the prospect of identifying super-enhancers in patient tumors and developing novel therapeutics to disrupt their control of key oncogenes," says Bradner.
Young meanwhile is increasingly confident that this super-enhancer paradigm of gene regulation has important implications across human disease states.
“Looking at large genome association studies, one can find disease-related mutations occurring in super-enhancers,” Young says. “It’s possible that super-enhancers could become biomarkers that identify key disease genes and help guide the development of approaches to treatment.
This work was supported by grants from the National Institutes of Health (grant HG002668), the National Cancer Institute (grant CA146445), and the Damon-Runyon Cancer Research Foundation.
***
Citations:
Whyte, W. A., Orlando, D. A., Hnisz, D., Abraham, B. J., Lin, C. Y., Kagey, M. H., ... & Young, R. A. (2013). Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell, 153(2), 307-319.
Lovén, J., Hoke, H. A., Lin, C. Y., Lau, A., Orlando, D. A., Vakoc, C. R., ... & Young, R. A. (2013). Selective inhibition of tumor oncogenes by disruption of super-enhancers. Cell, 153(2), 320-334.
Contact
Communications and Public Affairs
Phone: 617-452-4630
Email: newsroom@wi.mit.edu