Protein complex in key cell-growth pathway could help predict response to cancer therapy

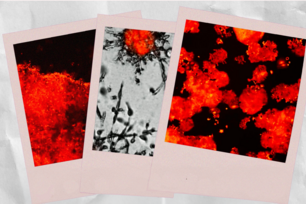
The protein complex GATOR1's location in cells where it is functional (left) and nonfunctional (right)
Liron Bar-Peled and Lynne Chantranupong/Whitehead Institute
CAMBRIDGE, Mass. – Whitehead Institute researchers have identified a protein complex that, when mutated, sends the master growth regulatory pathway known as mTORC1 into overdrive. Although mutations to the complex, dubbed GATOR1, are prevalent in cancer cells and promote tumor growth, they also render the cells susceptible to treatment with the drug rapamycin.
“We’re hopeful that genetic analysis of GATOR1 in tumors could help predict which cancer patients would respond to rapamycin, so there would be a clinical benefit to measuring these mutations,” says Whitehead Member David Sabatini.
When the mTORC1 (for “mechanistic target of rapamycin complex 1”) pathway operates properly, it senses the amount of nutrients, specifically amino acids, available and restricts cell growth based on that level. According to Sabatini, who is also a Howard Hughes Medical Institute investigator and professor of biology at MIT, this pathway is hyperactive in an estimated 50-80% of cancers, driving tumor cell growth and proliferation.
Rapamycin, an FDA-approved immunosuppressant used in organ transplants, targets the mTORC1 pathway, and may be able to interrupt the pathway’s hyperactivity in cancer cells. Yet researchers have been uncertain how to identify the tumors in which rapamycin might be effective.
In this week’s issue of the journal Science, researchers in the Sabatini lab describe how they identified this endogenous inhibitor (GATOR1 for “GAP activity toward Rags”) of the mTORC1 pathway. Liron Bar-Peled and Lynne Chantranupong, who are both authors of the Science article and graduate students in Sabatini’s lab, found that GATOR1 itself is mutated in several cancers, including glioblastomas and ovarian cancers, and that cancer cells with these mutations are also highly sensitive to treatment with rapamycin.
“We know that mTORC1 is largely deregulated in many, many different cancers,” says Bar-Peled. “A real problem in the field has been assigning good biomarkers, which would help to treat cancer patients with these pharmacological inhibitors, such as rapamycin. I think it’s really exciting that we showed that cancer cells deficient in these GATOR proteins are hypersensitive to this pharmacological inhibitor of mTOR, rapamycin. So for certain cancers, this could be really meaningful.”
But both he and Chantranupong caution that this research is still very preliminary.
“We need to investigate the physiological context of how rapamycin and GATOR1 operate in cancer cells,” says Chantranupong, suggesting that more work needs to be done in animal models before it can be translated to human patients. “Still it’s very promising and could apply to even more types of cancer than we’ve studied so far.”
This work was supported by grants from the NIH (CA103866 and AI47389), Department of Defense (W81XWH-07-0448), National Cancer Institute (NIH) (U24CA143867), David H. Koch Graduate Fellowship Fund; NSF Graduate Research Fellowship Program; Harvard-MIT Health, Sciences, and Technology IDEA2 program; and American Cancer Society.
* * *
David Sabatini's primary affiliation is with Whitehead Institute for Biomedical Research, where his laboratory is located and all his research is conducted. He is also a Howard Hughes Medical Institute investigator and a professor of biology at Massachusetts Institute of Technology.
* * *
Citation:
Bar-Peled, L., Chantranupong, L., Cherniack, A. D., Chen, W. W., Ottina, K. A., Grabiner, B. C., ... & Sabatini, D. M. (2013). A Tumor suppressor complex with GAP activity for the Rag GTPases that signal amino acid sufficiency to mTORC1. Science, 340(6136), 1100-1106.
Topics
Contact
Communications and Public Affairs
Phone: 617-452-4630
Email: newsroom@wi.mit.edu