Understanding the evolution of drug resistance points to novel strategy for developing better antimicrobials
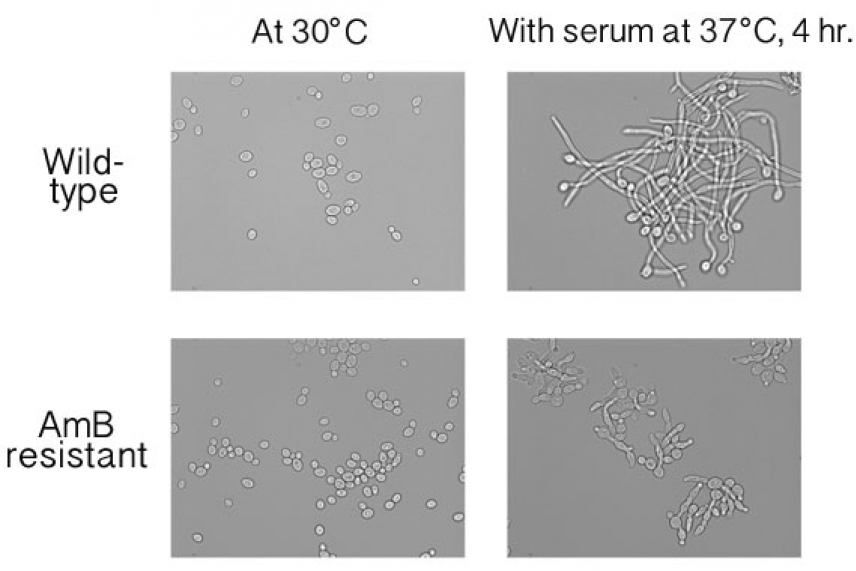
Filamentation in yeast with (top) and without amphotericin B resistance (bottom)
Benjamin Vincent/Whitehead Institute
CAMBRIDGE, Mass. – The rapid increase of drug-resistant bacteria, parasites, and fungi is fueling efforts not only to create new, more effective drugs, but also to understand the mechanisms behind the emergence of drug resistance. Rather than studying organisms known for tolerating antimicrobial drugs, Whitehead Institute scientists have come at the problem from the opposite perspective: by determining why one antifungal drug, amphotericin B, has remained refractory to drug resistance for five decades.
According to research by Whitehead Member Susan Lindquist’s lab, the remarkable resilience of amphotericin (AmB) against the emergence of resistance is due to the dire trade-offs between AmB-resistance and fungal virulence. Genetic mutations that enable certain strains of Candida albicans to resist to AmB also render the pathogen highly susceptible to the stresses of the host environment and immune system. AmB-resistant strains require high levels of the chaperone heat shock protein 90 (Hsp90) to survive such stress and are actually incapable of causing infection in a host. In addition, the mutations that confer AmB resistance in C. albicans disable the ability of the fungus to invade and penetrate mammalian tissue. Learning how AmB resistance limits C. albicans’ infectious capability provides insights for creating new antifungals and antibiotics that create high costs of resistance, thereby preventing the rise of resistant strains.
Drug resistance is an evolutionary process. As bacteria, fungi, viruses, and parasites are exposed to various antimicrobial agents, surviving strains became resistant to one or more drugs.
Much research to date has focused on understanding how drug-resistant organisms are able to dodge an onslaught of antimicrobials, with the ultimate goal of improving the next generation of pharmaceuticals. Scientists in Lindquist’s lab flipped this idea on its head, choosing instead to investigate why strains of C. albicans resistant to AmB are so rare. In fact, previous research has established that of more than 9,000 C. albicans strains sampled from patients, 99.8% remained susceptible to AmB.
C. albicans is normally a harmless commensal organism, but it can spiral out of control in immuno-compromised patients; it is the leading fungal pathogen in humans and the fourth-most commonly acquired hospital infection. C. albicans infections are generally treated with drugs from two major antifungal classes, triazoles and echinocandins. But the yeast can quickly evolve resistance to these drugs. Despite serious side effects including fever, tremors, and kidney toxicity, AmB has been used as a monotherapy for C. albicans for the past 50 years, a timespan more than sufficient to allow numerous resistant strains to emerge.
To understand why AmB resistance is so rare, the Lindquist lab investigated the mutations found in resistant clinical and laboratory strains. As reported in this week’s issue of the journal PLOS Biology, they found that even in the absence of AmB, resistant strains are highly dependent on Hsp90, a protein that helps other proteins function properly, enables cells to survive periods of stress, and plays a critical role in the evolution of new traits.
Cells normally have an overabundance of Hsp90 that acts as a “buffer” during stressful events. But AmB-resistant strains overload the demand for Hsp90. Cells are no longer able to respond to additional stressors they encounter when invading tissues and being attacked by the immune system. In fact, the pathogen is so compromised that mice injected with large numbers of AmB-resistant cells do not develop C. albicans infections.
Benjamin Vincent, a graduate student in Lindquist’s lab, says that knowing why C. albicans is virtually unable to become resistant to AmB may open up new ways to battle drug-resistance.
“This research indicates a different approach for the design of new antimicrobial agents,” says Vincent, who is also an author of the PLOS Biology paper. “Make a drug, like amphotericin B, that exploits the cost of resistance. The fungus or bacteria can’t evolve resistance to it without losing their ability to infect the host. We think that exploiting drug resistance to shut off virulence pathways in this way could be broadly applicable to other infectious diseases. And most directly, our work strongly supports the pursuit of new analogues of amphotericin with reduced toxicity to humans, an effort already underway in Martin Burke’s lab at University of Illinois.”
This work is supported by the National Science Foundation, Howard Hughes Medical Institute (HHMI), and the Mathers Foundation.
* * *
Susan Lindquist’s primary affiliation is with Whitehead Institute for Biomedical Research, where her laboratory is located and all her research is conducted. She is also a Howard Hughes Medical Institute investigator and a professor of biology at Massachusetts Institute of Technology.
* * *
Citation:
Vincent, B. M., Lancaster, A. K., Scherz-Shouval, R., Whitesell, L., & Lindquist, S. (2013). Fitness trade-offs restrict the evolution of resistance to amphotericin B. PLoS Biol, 11(10), e1001692.
Contact
Communications and Public Affairs
Phone: 617-452-4630
Email: newsroom@wi.mit.edu

