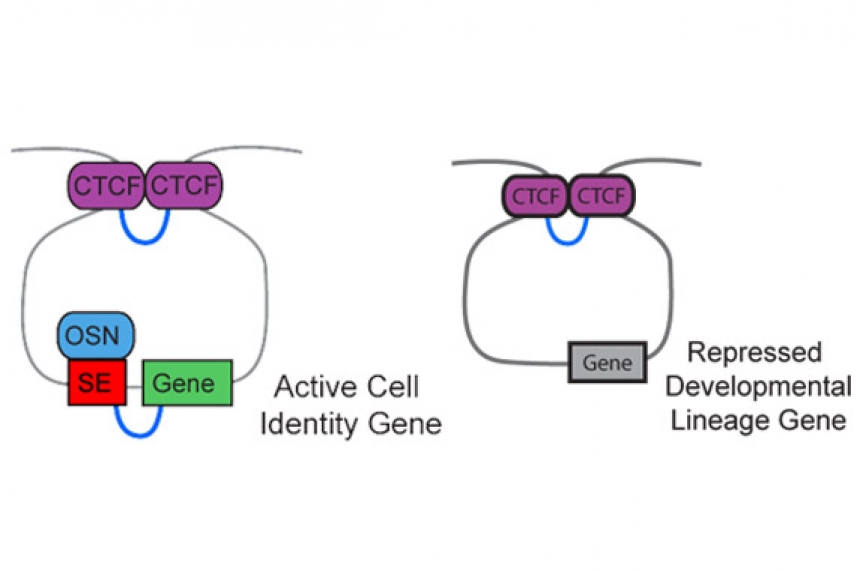
The proteins Cohesin (blue lines) and CTCF (purple blocks) act as the purse strings of DNA "goody bags" (gray lines). The bags' locations in the genome are similar across cell types, although the bags' contents and roles can change from transcription factors (blue OSN oval) and enhancers (red box) that promote the gene (green) expression (left diagram) or to repressors (right diagram) that inhbit gene expression (gray box).
Courtesy of Cell Press
Special chromosomal structures control key genes
CAMBRIDGE, Mass. – Within almost every human cell is a nucleus six microns in diameter—about one 300th of a human hair’s width—that is filled with roughly three meters of DNA. As the instructions for all cell processes, the DNA must be accessible to the cell’s transcription machinery yet be compressed tightly enough to fit inside the nucleus. Scientists have long theorized that the way DNA is packaged affects gene expression. Whitehead Institute researchers present the first evidence that DNA scaffolding is responsible for enhancing and repressing gene expression.
“For the first time, we see that the structure of the chromosomes contributes to gene control,” says Whitehead Member Richard Young, who is also a professor of biology at MIT. “In the past, there have been all kinds of ideas around how the structure might affect gene control, but now one has been experimentally tested and shown to be true.”
DNA scaffolding has a hierarchy that ranges from beads-on-a-string, where a strand of DNA is wrapped around histone proteins to form a nucleosome, to topological associating domains akin to DNA balls containing multiple DNA loops interacting with various regulatory elements, to highly condensed chromosomes.
Working in mouse embryonic stem cells (ESCs), the Young lab examined the scaffolding landscape found within topologically associated domains. The researchers knew that proteins called Cohesin and CTCF are bound to the DNA around important ESC genes, but they did not know how these proteins affect the DNA’s three-dimensional organization. Using a technique known as ChIA-PET, the researchers focused on how these proteins interact.
“By knowing which of the Cohesin/CTCF bound sites are coming together in physical proximity, we started to go from a linear view of the genome to sets of looping interactions, which led us to these domains, these super enhancer domains, where gene expression enhancement is contained within the loop,” says Jill Dowen, a postdoctoral researcher in Young’s lab.
Dowen and coauthors Zi Peng Fan, Denes Hnisz, and Gang Ren describe the structure of these loops in the latest online edition of the journal Cell.
Hnisz, also a postdoctoral researcher in Young’s lab, likens the loops to “goody bags”, with Cohesin and CTCF acting as the purse strings to create a DNA loop that cradles proteins enhancing or repressing gene expression. The enhancement or repression effects are confined to the genes within the goody bag. In the ESCs they studied, the scientists identified 197 Cohesin/CTCF-flanked loops that contain active genes and enhancers, and 349 loops that contain repressed genes.
Interestingly, when mouse neural precursor cells were examined for similar Cohesin/CTCF-bound loops, not only did the cells have such loops, but they were in similar locations as in the ESC genome. Of course, the repression or enhancement role of the loops differed between ESCs and the more differentiated cells. This indicates that the loops may remain constant during a cell’s differentiation, although they may switch their contents, and therefore their effect on gene expression.
The researchers plan to study the loop structure and placement in other types of differentiated cells, but for now, they are excited about their new appreciation of DNA structure and its effects.
“I think we’ve filled in a major gap in the understanding of how gene expression is linked to the local organization of chromosomes,” says Hnisz. “It will be exciting to explore whether defects in these mechanisms might even contribute to human disease.”
This work is supported by National Institutes of Health (NIH, HG002668, CA168263-01A1), Austrian Science Fund (FWF, J3490), and Netherlands Organization for Scientific Research (NWO). Young is a founder of Syros Pharmaceuticals.
* * *
Richard Young’s primary affiliation is with Whitehead Institute for Biomedical Research, where his laboratory is located and all his research is conducted. He is also a professor of biology at Massachusetts Institute of Technology.
* * *
Citation:
Dowen, J., Fan, Z., Hnisz, D., Ren, G., Abraham, B., Zhang, L., . . . Young, R. (2014). Control of Cell Identity Genes Occurs in Insulated Neighborhoods in Mammalian Chromosomes. Cell, 159(2), 374-387. doi:10.1016/j.cell.2014.09.030
Contact
Communications and Public Affairs
Phone: 617-452-4630
Email: newsroom@wi.mit.edu


