Paired genes in stem cells shed new light on gene organization and regulation
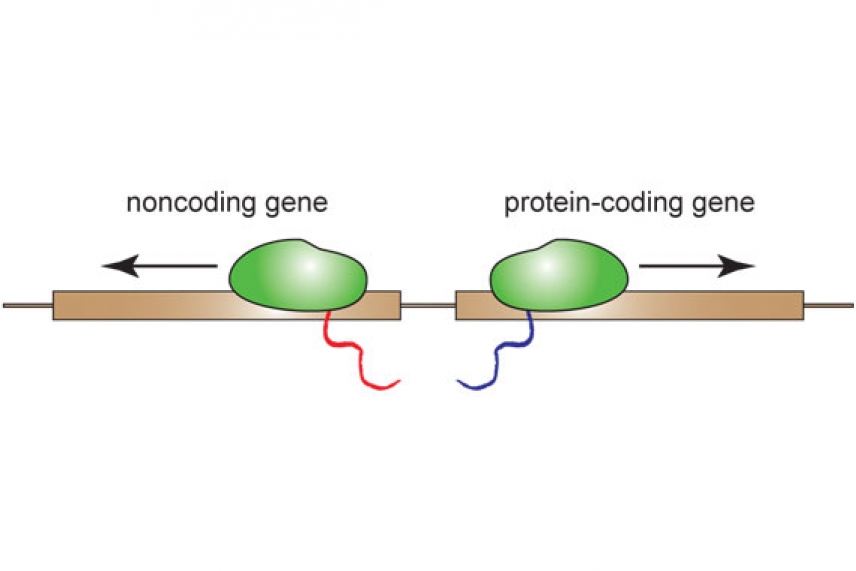
Diagrm of transcription of mRNA and lncRNA that start at the same location
Alla Sigova
CAMBRIDGE, Mass. – Whitehead Institute researchers have determined that DNA transcription, the process that produces messenger RNA (mRNA) templates used in protein production, also runs in the opposite direction along the DNA to create corresponding long noncoding RNAs (lncRNAs). Moreover, the mRNAs and lncRNAs are transcribed coordinately as stem cells differentiate into other cell types. This surprising finding could redefine our understanding of gene organization and its regulation.
“It’s a surprise to me that genes come in pairs,” says Whitehead Member Richard Young, who is also a professor of biology at MIT. “At any one of the 20,000 protein-coding genes that are active in human stem cells, a lncRNA gene located upstream is also transcribed. So much effort has gone into studying protein-coding genes, and yet we have missed this concept that all protein-coding genes come in mRNA/lncRNA pairs. If you activate the mRNA gene, you’re going to activate the lncRNA gene.”
Young and his lab report their findings this week in the Proceedings of the National Academy of Sciences (PNAS).
Until now, scientists thought transcription machinery attaches to DNA at certain points called promoters and moves just in one direction along the DNA to create mRNAs from protein-coding genes. Other RNAs that are not protein templates, including lncRNAs, are also created by transcription, but despite their important roles in in regulation of gene expression, development and disease, scientists knew little about how lncRNA transcription is initiated or where most lncRNA genes reside in the genome.
By looking at human and mouse embryonic stem cells, researchers in the Young lab found something astonishing—most lncRNA genes are located adjacent mRNA genes, and the transcription of about 65% of lncRNAs originates at active promoters associated with these mRNAs’ genes and runs “upstream” and in the opposite direction from the promoter.
When the transcription machinery attaches to a promoter, it seems that it is just as likely to move in one direction and transcribe the mRNA as it is to move in opposite direction and transcribe the neighboring lncRNA.
The researchers also noticed that as embryonic stem cells begin differentiating into other cell types, the mRNA/lncRNA pairs are regulated in the same way—the transcription of paired mRNAs and lncRNAs is upregulated or downregulated together. This further confirms the relationship between the transcription of mRNAs and lncRNAs.
“I think it’s quite a breakthrough,” says Alla Sigova, who is a postdoctoral researcher in the Young lab and a co-author of the paper in PNAS. “This provides a unifying principle for production of mRNAs and lncRNAs, and may lead to new insights into lncRNA misregulation in disease. For example, some mRNAs are up- or downregulated in cancer, so their lncRNA partners could potentially contribute to cancer.”
This work was supported by National Institutes of Health Grants HG002668, GM34277, and DK090122, the American Gastroenterological Association, the Damon Runyon Cancer Research Foundation, and National Cancer Institute Cancer Center Support (Core) Grant P30-CA14051.
* * *
Richard Young's primary affiliation is with Whitehead Institute for Biomedical Research, where his laboratory is located and all his research is conducted. He is also a professor of biology at Massachusetts Institute of Technology.
* * *
Citation:
Sigova, A. A., Mullen, A. C., Molinie, B., Gupta, S., Orlando, D. A., Guenther, M. G., ... & Young, R. A. (2013). Divergent transcription of long noncoding RNA/mRNA gene pairs in embryonic stem cells. Proceedings of the National Academy of Sciences, 110(8), 2876-2881.
Topics
Contact
Communications and Public Affairs
Phone: 617-452-4630
Email: newsroom@wi.mit.edu


